A recent study of a type of immune blood cells associated with resistance to certain treatments for melanoma is one sign of the growing role of data science in solving some of medicine’s most puzzling riddles, says Hatim Sabaawy, MD, PhD, associate director of translational research at the University of Colorado Cancer Center.
“I have a son who is still an undergrad, and I told him, ‘You can be going to top places with training in bioinformatics and improving your computer skills,’” says Sabaawy, professor in the Division of Medical Oncology of the CU Department of Medicine.
A research project like this and others like it require a multi-disciplinary approach involving people with varied backgrounds, from cancer biology to computer science, he says. It’s the kind of research approach that has become a top priority at the CU Cancer Center, not just for fighting melanoma but other cancers and other diseases as well.
“In the last five years or so, there has been an exponential growth in these types of studies where we're looking at how tumor micro-environments differ in terms of responses to therapy, and in discovery of new biomarkers that could be used for developing new types of therapies,” Sabaawy says. “And not just one or two or three types of markers at a time. Now we can do large-scale studies on hundreds or thousands of markers.”
A powerful yet tiny image
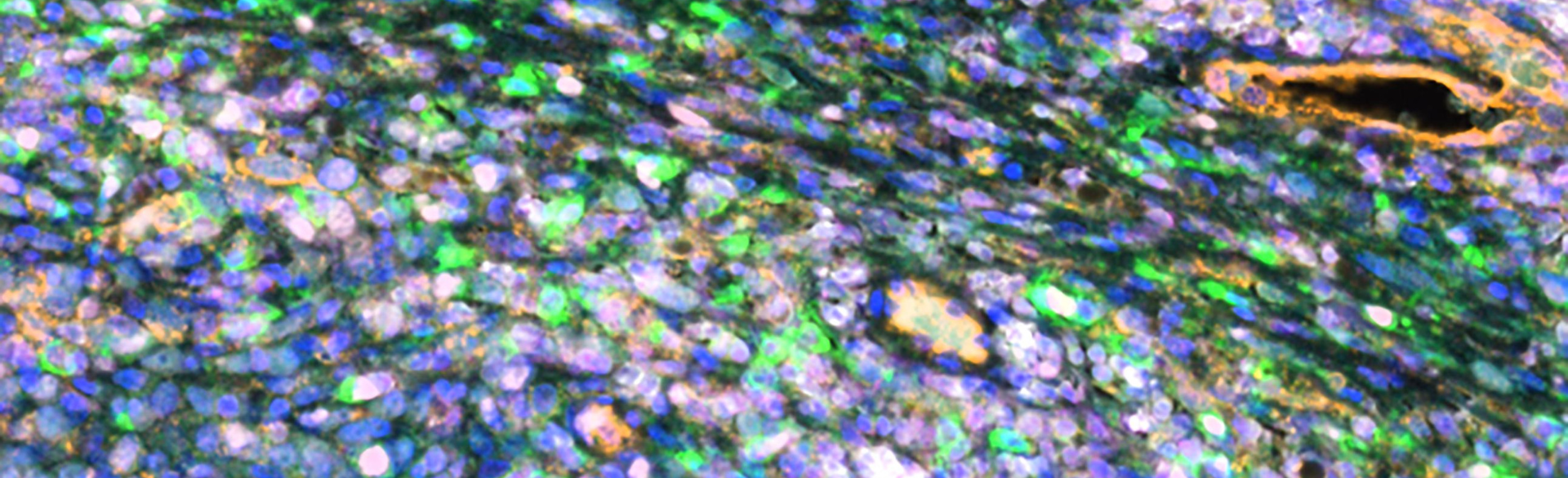
One dazzling example of data analysis as a tool of cutting-edge research is the colorful image that appears on the cover of the Oct. 15 issue of the journal Clinical Cancer Research, in which an article on the recent melanoma study was published. The image – encompassing a tiny area no bigger than a house dust mite – shows part of a patient’s melanoma tumor and surrounding cells.
Ann Strange, MBA, a bioinformatics developer in Sabaawy’s lab, helped create the image and spearheaded image analysis on the research project, which she co-authored.
The first step in creating the image, she says, was to obtain slides of specially prepared patient biopsy samples from the School of Medicine’s Melanoma Biorepository. Those slides were then processed by the Human Immune Monitoring Shared Resource (HIMSR), a center at the CU Cancer Center that provides technical support for sophisticated immune-response analysis. Colors in the image were digitally separated to make different cells being studied more visible – the melanoma tumor itself as well as parts of cells.
The image helped researchers differentiate types of cells and cell markers and to figure out how tumor cells are interacting with surrounding cells. This kind of imagery in a data science approach called spatial profiling has only become possible in recent years as powerful computing, multiplex fluorescence imaging and sophisticated software analysis codes have become available.
Strange, who previously worked for financial and tech companies, comes from a computer science background. “I’m thrilled that I can apply those skills to a biomedical application,” she says. “It definitely helps to have the coding and machine-learning tools available to produce these data in an automated way. It's becoming ever more complex. And in this case, we looked for an image that helps characterize and add further color to findings from other data.”
Targeting therapies
David Woods, PhD, led the study while an assistant professor at the CU Cancer Center. Woods recently left the university to start a biotech company but is wrapping up work on CU Cancer Center projects.
The heavily data-driven study comes as more immunotherapies – treatments designed to manipulate the immune system to fight melanoma and other cancers – are being introduced. “But not all patients do well” under such treatments, Woods says, “and the big question in the field is, why do some patients do well and some don’t? And can we identify which patients will do well and which will not, and then choose effective treatment strategies based on that?”
The new study focused on T-cells, which are a type of immune blood cells that normally lead the body’s fight against disease. But as previous studies by Woods, Strange and their colleagues showed, elevated levels of certain types of T-cells in melanoma patients’ blood are associated with resistance to a variety of immunotherapy drugs called checkpoint inhibitors. These immunotherapies are designed to deactivate a biological off switch that might cause T-cells to stop fighting cancer in the body because they mistake cancer cells for healthy tissue.
The new study found an association between checkpoint-inhibitor resistance and a type of T-cells called CD4+ that produce certain enzyme markers labeled CD38 and CD39. Those cells are found in the blood of some melanoma patients and in the tumors themselves.
“It’s not just that the T-cells no longer kill tumors,” Woods says. “They’re actually switching sides and suppressing the immune response.”
Potential for follow-up
The study concludes that these T-cells are “a potential rational target to improve the efficacy of checkpoint inhibition.”
“There is a lot of potential for follow-up on these types of studies here at the CU Cancer Center,” Sabaawy says. Using data in ever more sophisticated ways to unlock cancer mysteries “is the approach that we’re putting a lot of effort into at the CU Cancer Center, so having people with Ann’s expertise is becoming more and more important,” he says.
“At the medical school, we are improving the way we incorporate data science into the curriculum,” he adds. Also, the CU Cancer Center is working with the new CU Department of Biomedical Informatics (DBMI) on investigating “how data science analysis can be incorporated into translational and clinical studies from the start.” The DBMI launched in 2022.
Image at top courtesy of Ann Strange. It shows a section of a tumor biopsy from a patient with metastatic melanoma. Immunofluorescence staining shows infiltration of CD4+ T-cells expressing the enzyme markers CD38 and CD39.



.png)